Novel human NFKB2 protein mutation in two unrelated patients with common variable immunodeficiency
What is common variable immunodeficiency?
Common variable immunodeficiency (CVID) is one of the most common, clinically significant primary immunodeficiencies(1). It is caused due to genetic defects that result in impaired B-cell differentiation leading to defective antibody production. Mutations in approximately 23 different genes have been linked to CVID, many of which are involved in B-cell differentiation and antibody production(2). Most patients have sporadic disease, but 10–25% of them have familial inheritance. Genetic testing is therefore a key aspect of CVID diagnosis. Respiratory tract infections are the most common among patients due to the absence of protective antibodies3. Nevertheless, there are also patient subgroups that present with a range of other complications including malignancy, immune dysregulation and autoimmune conditions(4).
What is the link between CVID and nuclear factor kappa-B subunit 2?
Evidence from published cases indicates that approximately 5% of the CVID population has mutations in the nuclear factor kappa-B subunit 2 (NFKB2) gene that encodes the p100 subunit of the transcription factor complex NFKB and is a key player in the non-canonical NFκB signalling pathway(2).
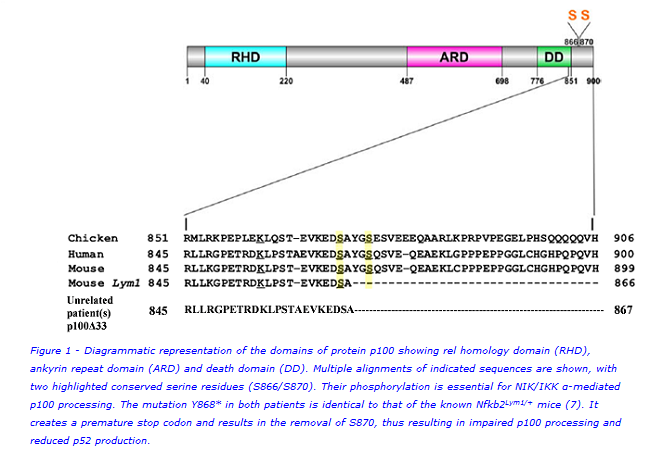
Figure 1 is a diagrammatic representation of the domains of NFKB2 (p100) showing the rel homology domain (RHD), ankyrin repeat domain (ARD) and death domain (DD). Multiple alignments of indicated sequences are shown, with two highlighted conserved serine residues (S866/S870). When the non-canonical NFκB pathway is activated, these serines are phosphorylated which marks the p100 protein for partial cleavage into functional p52 that then translocates to the nucleus of the cell to initiate transcriptional regulation of downstream target genes(5).
Clinical Implications
Lack of antibodies, recurrent infections, adrenocorticotropic hormone (ACTH) deficiency and autoimmune hair loss are features in patients that have been associated with mutations in the NFKB2 gene(2,6). We therefore postulated a causative NFKB2 gene defect in two unrelated cases from Epsom & St. Helier University Hospitals, with these clinical features. To verify this, we sequenced the NFKB2 gene in these patients, measured the expression of protein p100, evaluated B- and T-cell subsets and performed quantitative functional assessment of the non-canonical NFκB signalling pathway.
Sequencing identified novel heterozygous genetic variants at position 2604 in the NFKB2 gene in both patients, causing the same amino acid change i.e tyrosine to termination at position 868 in the encoded protein p100. This resulted in a truncated form of p100, p100Δ33, which is 33 amino acids shorter than the normal form of the protein. Interestingly, this mutation in the two unrelated patients is identical to that of the known Nfkb2Lym1/+ mice(7). Loss of serine870 disrupts the phosphorylation essential for processing of p100 to p52. Immunoblotting showed accumulated unprocessed mutant p100Δ33 and reduced p52. DNA binding by functional p52 was quantitatively abrogated. Circulating B and T cells (including T follicular helpers) were significantly reduced.
Thus, patients presenting with lack of antibodies, recurrent infections, ACTH deficiency and autoimmune hair loss should be tested for mutations in the NFKB2 gene. Further research using samples from these unrelated patients and the Nfkb2Lym1/+ mice will help us better understand the molecular pathogenesis of CVID.
This study was undertaken by
Kruthika Sundaram(a,b), John Guly(c), Susan Tadros(c), Frances Smitha(a,b), Terry Hunter(a,b), Helene Martini(a), Ioasaf Karafotias(a), Grant Hayman(c), Mohammad A A Ibrahim(a)
a. King’s College London, King’s Health Partners, King’s College Hospital NHS Foundation Trust, School of Medicine, Division of Asthma, Allergy & Lung Biology, Department of Immunological Medicine, Denmark Hill, London, UK
b. Viapath, King’s College Hospital, Denmark Hill, London, UK
c. Epsom & St Helier University Hospitals NHS Trust, Carshalton, UK
For further information, please contact:
Dr. Kruthika Sundaram:
kruthika.sundaram [at] nhs.net
References:
- Durandy A, Kracker S, Fischer A. Primary antibody deficiencies. Nat Rev Immunol. 2013;13(7):519-533.
- Bogaert DJ, Dullaers M, Lambrecht BN, Vermaelen KY, De Baere E, Haerynck F. Genes associated with common variable immunodeficiency: one diagnosis to rule them all? J Med Genet. 2016;53(9):575-590.
- Bonilla FA, Geha RS. Common variable immunodeficiency. Pediatric Research. 2009;65(5):13–9.
- Gathmann B, Mahlaoui N, et al. for the ESID Registry Working Party. Clinical picture and treatment of 2212 patients with common variable immunodeficiency. Journal of Allergy and Clinical Immunology. 2014; 134(1):116–126.
- Sun SC. Non-canonical NF-kappaB signaling pathway. Cell Res. 2011;21(1):71-85.
- Liu Y, Hanson S, Gurugama P, Jones A, Clark B, Ibrahim MA. Novel NFKB2 mutation in early-onset CVID. J Clin Immunol. 2014;34(6):686-690.
- Tucker E, O’Donnell K, Fuchsberger M, et al. A novel mutation in the Nfkb2 gene generates an NF-kappa B2 “super repressor”. J Immunol. 2007;179(11):7514-7522

